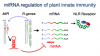
Our research focuses on understanding molecular-genetic mechanisms of plant innate immunity. We are investigating the structure, function and evolution of host genes for pathogen disease resistance. Our experimental system includes viral, bacterial and oomycete plant pathogens and their Solanaceae plant hosts. We anticipate that our studies will lead to new environmentally benign strategies for durable, broad-spectrum disease resistant crops.
Tobacco mosaic virus (TMV) triggers the N (Necrotic) resistance gene-mediated hypersensitive response (HR, brown lesions).